By Rosanna Tofalo
“Vino cotto” is a sweet wine, produced according to traditional procedures involving a prolonged fermentation of cooked grape must. It is inserted in the Italian list of traditional food products since 2000 and can be marketed as traditional agrifood product. The flow-chart for the traditional manufacture of “vino cotto” is showed in figure 1. Cooked must is produced from white or red grapes of the cultivars Trebbiano, Passerina, Moscato, Montonico, Sangiovese, and Montepulciano. It is obtained by direct heating of the must in copper boilers until its volume is reduced by 30–70%, according to the specific process. During cooking, the must becomes dark and dense, and reaches a very high sugar concentration (up to 55% w/v).
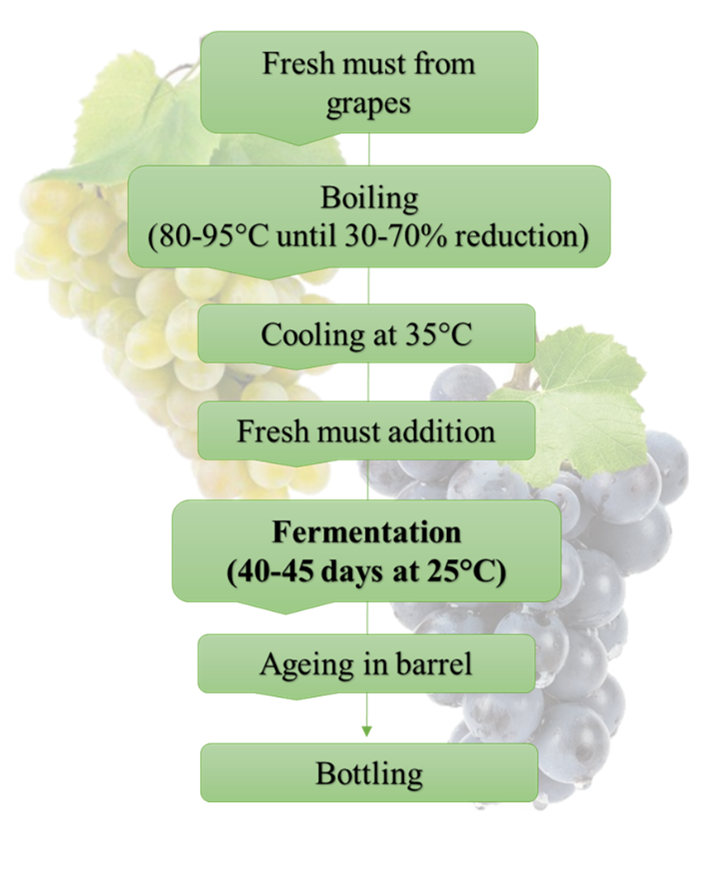

To start alcoholic fermentation, fresh must is added to sterile cooked must and the indigenous yeasts can drive the fermentation, which proceeds slowly (about 40 days at room temperature). Stuck or sluggish fermentations may occur during the production of “vino cotto”. These problems derive from the high osmotic conditions of cooked must which brings to hyperosmotic shock, followed by cytoskeleton collapse, intracellular damage, and subsequent arrest of cells growth. The majority of yeasts requires a minimum water activity (aw) of 0.85 for their growth, and only xerophilic yeasts (aw 0.61 and 0.75) can grow. Only few species can be detected in “vino cotto” wine samples: Saccharomyces cerevisiae, Candida apicola, Starmerella bacillaris (syn. Candida zemplinina) and Zygosaccharomyces bailii. All the species present osmotolerant traits being able to develop in presence of high concentration of glucose in a strain dependent way.
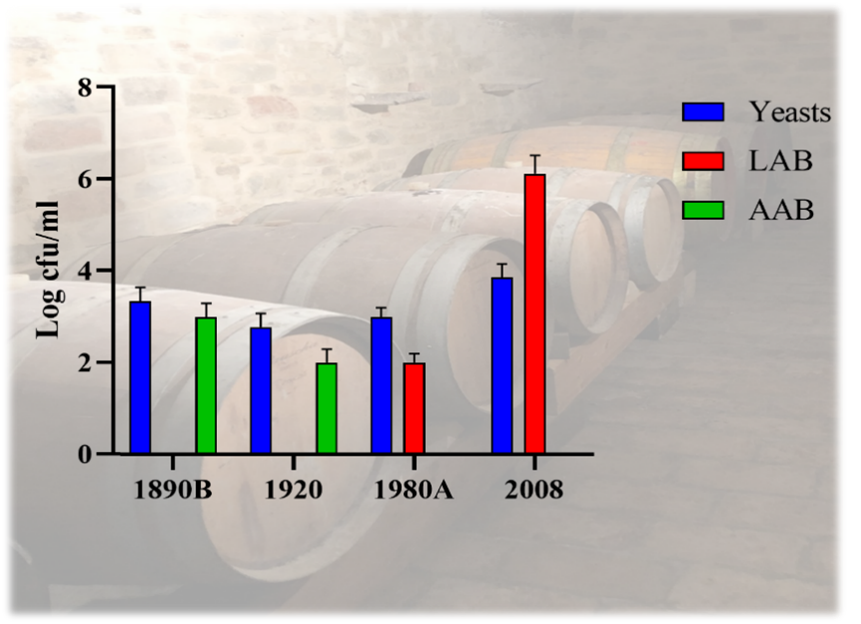
Recently we determined the cultivable microorganisms associated with “mother” of “vino cotto” collected from barrels of different years (1890, 1920, 1980 and 2008) (Figure 1). “Mother” of “vino cotto” is a mixture of must/wine settled in the bottom of barrels. It is a highly selective environment. Viable count of yeast communities showed values ranging from 2.7 Log cfu/mL (1890) to 3.5 Log cfu/mL (2008). Yeast counts increased over time showing the highest values in samples obtained from the newest barrels. Lactic acid bacteria (LAB) showed a similar trend. They were present only in “mothers” of “vino cotto” collected from the most recent barrels ranging from 2 Log cfu/mL (1980) to 6.12 Log cfu/mL (2008). Acetic acid bacteria (AAB) were detected in the oldest 3 samples with values from 4 Log cfu/mL (1890) to 2 Log cfu/mL (1920), while were absent in “mothers” of “vino cotto” obtained from the newest barrels (1980, 2008). Therefore, samples from barrels of 1890, and 1920 were characterized by the presence of yeasts and AAB, while those of 1980 and 2008 by yeasts and LAB (Figure 2). In particular, bacteria isolates belonged to LAB (Lactiplantibacillus plantarum and Pediococcus pentosaceus) and AAB (Gluconobacter oxydans). Starmerella lactis-condensi, Starm. bacillaris, Hanseniaspora uvarum, S. cerevisiae, Hanseniaspora guillermondi and Metschnikowia pulcherrima were the yeast species detected.
The source of a strain/species is a key factor affecting the final genetic diversity of the individual population. The identified microorganisms play a major role in the definition of final product characteristics, and they represent a core microbiota conserved over years. A better knowledge of the resident population present in the barrels can be exploited for the selection of ad hoc strains for “vino cotto”.

Rosanna Tofalo, obtained her PhD degree in Food Biotechnology at the University of Basilicata (2003). Since 2015, she is an associate professor at the Faculty of Bioscience and Technology for Food, Agriculture and Environment (University of Teramo, Italy). She teaches on food, industrial and wine microbiology. She is interested in the optimization of molecular methods for the detection, quantification and typing of food microorganisms. Furthermore, she carries out her research on microbial physiology, on yeasts and bacteria metabolism such as the production of secondary compounds during the fermentation process or enzymatic activities in order to select strains for biotechnological applications.
Email contact: rtofalo@unite.it

